Challenge Mechanics
The phases of the challenge
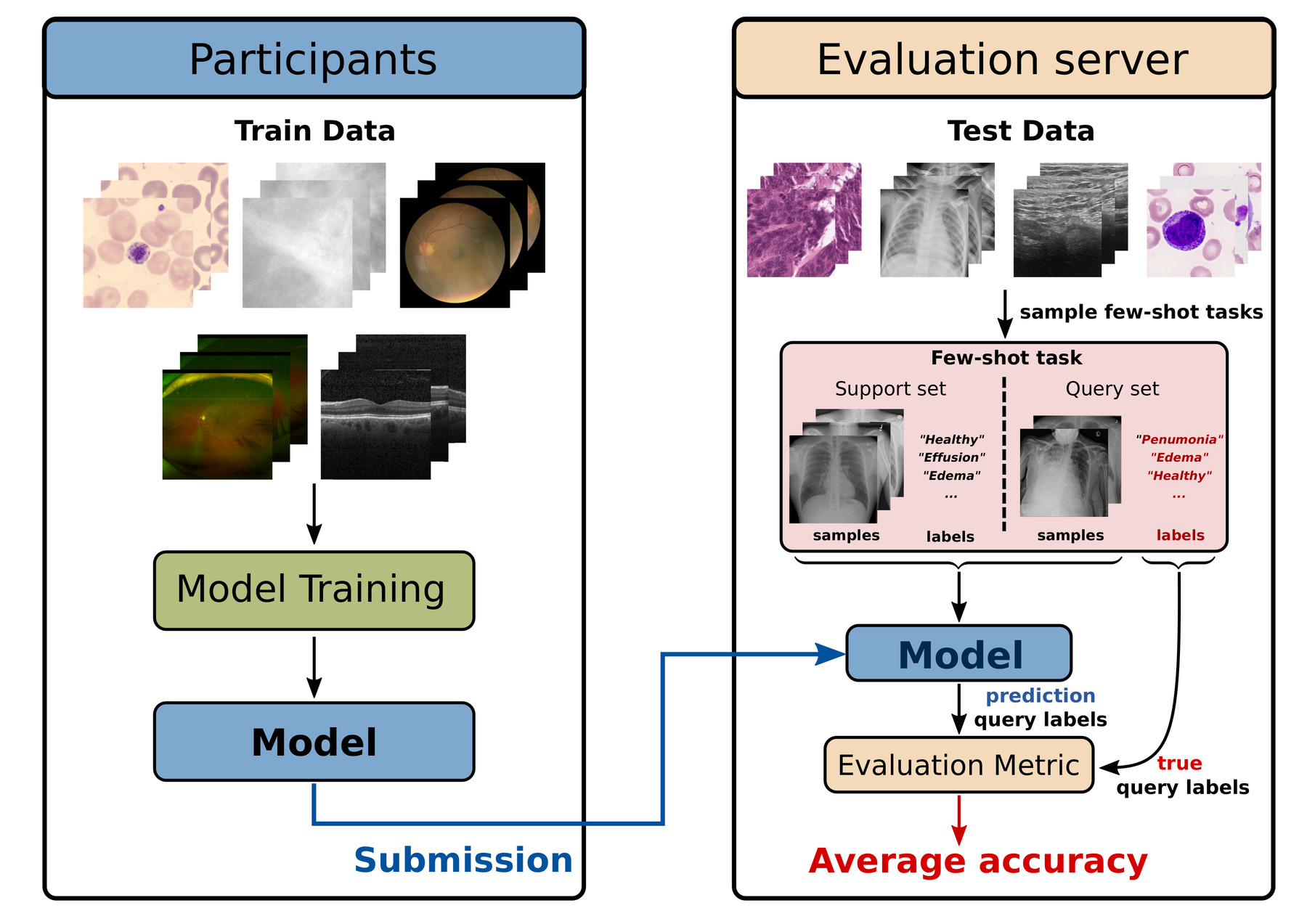
An overview of the submission and evaluation procedure are shown in the figure above. For a more detailed explanation checkout our Rules page.
Development and training
- We provide a meta-train dataset (i.e. a dataset consisting of multiple datasets) with multiple imaging modalities varying widely in type, e.g. binary, multi-class, or multi-label classification.
- Participating teams train a cross-domain few-shot learning algorithm with the provided data, and locally assess its performance by checking its ability to learn a left-out task using a small number of N of examples. Code examples and baselines will be provided.
- Ideas for algorithmic approaches can be drawn for example from meta-learning, transfer learning, or self-supervised pretraining.
Online validation
- When the submission platform opens (planned on 01 June 2023) algorithms can be submitted to the challenge server for validation on a private set of validation tasks. The algorithm should be packaged as a Singularity or Docker container and should accept the following inputs: a labelled support set, and a unlabelled query set for which predictions should be obtained. Boiler plate code and examples for building a Singularity or Docker container and a valid submission will be provided together with the opening of the submission platform.
- Validation submissions serve to allow participants to test the validity of their algorithms and will be used to populate an online leaderboard. However, the validation phase is distinct from the final test phase, and does not influence the final ranking in any way.
Final submission
- Until the submission deadline (planned 27 July 2023) participants are allowed to make 3 final submissions. Final submissions are made as a Singularity or Docker container exactly as in the validation phase. We will evaluate all submissions and use the best of the three for the final ranking.
- Note that teams planning to submit an algorithm must register before 22 June 2023.
- The final submission will be evaluated on a number of secret test tasks derived from secret test datasets that are distinct from the training and validation data. (Note that the images shown in the figure above are just for illustration purposes and are not from the real test data). The test data will include some data domains that are similar to some of the training data, and some data domains that are completely new.
- The winners will be announced at the MICCAI conference.
Evaluation
Metric
- On the evaluation server, we evaluate the algorithms on a set of hidden test tasks. For each task we create 100 few-shot learning task instances by randomly sampling a small support set of N labelled examples per class, along with a corresponding query set. The submitted algorithms should (on the challenge server) learn the new task, and output predictions for the query set. The accuracy on the query set will be our evaluation metric.
- We will evaluate all algorithms for a range of N from 3 to 10 shots for each test task. The average performance over all N in that range will be used as the final task performance.
Final ranking
- The average accuracies above are used to generate rankings for each test task. These rankings are averaged to generate a final aggregate challenge ranking. We use this rank-then-aggregate approach to not implicitly weigh any tasks higher than others.
